CRISPR/Cas9 gene editing for Zebrafish is a rapidly growing molecular biology toolset with diverse applications, but is not without its technological barriers. Reagent sourcing, design and validation of cutting efficiency, and donor homology design are challenging, lengthy processes without guarantee of success even for highly skilled researchers.
One way to enable researchers to initiate a project with confidence in the reagents they’ve selected is to rigorously test their sgRNA site. In fact we have found that validating sgRNA cutting efficiency is a critical step to mitigate failure rates in everything from null generation to homology driven repair and precise knock in.
Based on our conversations with customers, a failed zebrafish CRISPR knock-in project can cost a lab 9-12 months and an estimated $10K. But between lost labor and reagents, the real cost to a lab is usually much higher, particularly when multiple attempts are made to move a project forward. Researchers usually think the failure is further down in the process, when in fact, poor sgRNA cutting efficiency is often to blame. So in our labs, we consider validation of our sgRNAs to be one of the most important steps before moving forward with any CRISPR editing project.
sgRNA validation is not as straightforward as one might think. It requires expert knowledge, an experienced team, and a well-established system for interrogating cutting efficiency. The current DIY pathway for designing and verifying a selected sgRNA normally takes multiple weeks and may include the following steps:
- Identify the region of the genome that you want to modify
- Design using a variety of online algorithms to find potential sgRNA sites
- Identify predicted sgRNA sequences that have high cutting efficiency, but a low probability of off-target cutting.
- Order sgRNA sequence
- Clone sgRNA sequence into an expression vector
- Confirm you’ve cloned your sgRNA correctly using sequencing
- Synthesize and purify single stranded sgRNA
- Perform test Injections
- Select a method for verifying cuts
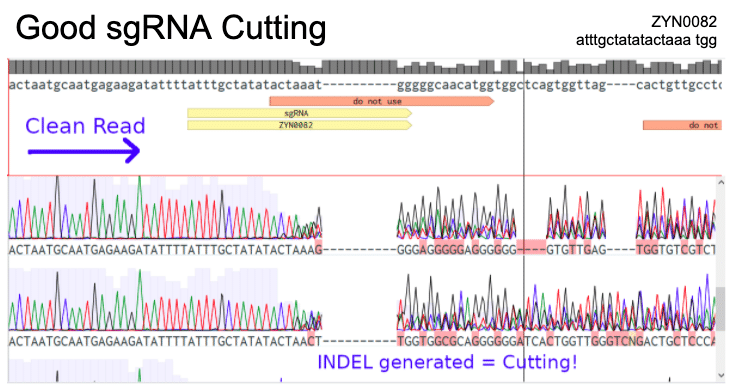
- Verify cutting efficiency
Importantly, cutting efficiency needs to be confirmed in vivo, not just in silico or in vitro. Currently, researchers use a variety of online algorithms to identify the best sgRNA sequence to for their project. Even sgRNA sequences with the highest scores from these algorithms are at risk for cutting inefficiently, because guided cutting efficiency of an sgRNA can be impacted by chromatin wrapping and genomic organization around your target site.
In addition, potential polymorphisms and SNPs in your locus need to be taken into account. Here differences between laboratory strains and published genomic assemblies can significantly impact the cutting efficiency of a selected sgRNA.
Until this verification process is completed, these risks will impact the workflow. Furthermore, even when workflows move smoothly, certain sgRNA sequences can be lethal when targeting a specific region of the genome. This means researchers may need to return to square one and repeat the entire process of selection and verification should any issues arise with the sgRNA they’ve selected.
Many researchers choose to use customized injection mix due to budget constraints. While this approach is great for researchers with significant in-house screening experience, we strongly recommend that customers also use our in vivo sgRNA verification process to improve the success rate and ensure no valuable time and energy are wasted along the way.

Marnie is a Staff Scientist and leads the Zebrafish Team at InVivo Biosystems, specializing in genome editing with CRISPR/Cas9 technology in zebrafish. She received her PhD in Neuroscience at Oregon Health and Sciences University in Portland in 2011, and completed her postdoctoral training in Cell & Developmental Neurobiology at the University of Oregon in Eugene, OR, and then at the University of Colorado Anschutz Medical Campus in Aurora, CO, where she worked to develop whole animal screening models for preclinical drug screening. Since joining InVivo Biosystems in 2018, she has enjoyed helping clients meet their research goals by generating custom zebrafish. In her spare time she enjoys exploring the beauty of the Pacific Northwest, cooking overly complicated meals and talking zebrafish with anyone who will listen