Crispants: Fast and Efficient Zebrafish Genetic Model Creation
Transform Your Research with Our Accelerated and
Efficient Gene Knockout Solution
Rapid Zebrafish Genetic Model Creation Using Crispants
More Zebrafish CRO Services
What is Crispant?
Crispants are a groundbreaking advancement in CRISPR/Cas9 technology. They empower researchers with a swift and cost-effective method description to produce F0 mutant alleles in zebrafish. Here are the notable benefits:
- Speed:
- From gene disruption to observing behavioral phenotypes, the Crispant method drastically reduces the usual timeline from over six months to just a week.
- Efficiency:
- This innovative approach minimizes animal use and offers an efficient strategy for disease modeling, proof-of-concept research, and targeted double-strand breaks for prime target validation.
- Cost-Effective:
By accelerating the research process, Crispants helps in significantly cutting down costs.
For researchers keen on screening loss-of-function alleles in vivo, Crispants are not just a tool but a game-changer, reshaping standard genetic approaches.
The InVivo's Crispants Difference
While traditional CRISPR techniques might take nearly a year, our proprietary Crispant approach fast-tracks KO knock-in lines:
Rapid Results: Generate transient knock-out lines in under a month.
Stable Knock-out: Use these transient zebrafish models to create long-term stable zebrafish knock-out models.
Benefits of Partnering with InVivo
Accelerated Decisions: Determine the viability of a functional genetic model in vivo in less than a month.
Simultaneous Assessments: Evaluate multiple gene functions in a shortened timeframe.
Dual Advantage: Start on full model creation while simultaneously gathering preliminary data.
Ideal Use Cases for Our Crispant Technology
Gene Exploration: Examine the biological function of individual genes or combinations.
Lethality Analysis: Assess the formation of loss of genes.
Preliminary Screening: Determine the necessity of creating a stable custom mutation.
Target Evaluation: Swiftly analyze potential targets for therapeutic solutions.
Specific Gene Targeting: Focus on particular functional regions within your candidate gene of interest.
What We Offer
Crispant Verified Injection Mix: A ready-to-inject mix that contains 3 sgs and the Cas9 enzyme. We identify and test 5 sgs across your gene, selecting the top three for the best in vivo editing efficiencies.
Crispant In Silico Predicted Injection Mix: Another inject-ready mix that includes 3 sgs and the Cas9 enzyme, chosen based on predictive editing efficiency score.
See the Efficiency of Crispant in Action
Replicating Heartstrings(tbx5a) mutant using crispant technique. Achieve knock-out in under a month with our Crispant technique.
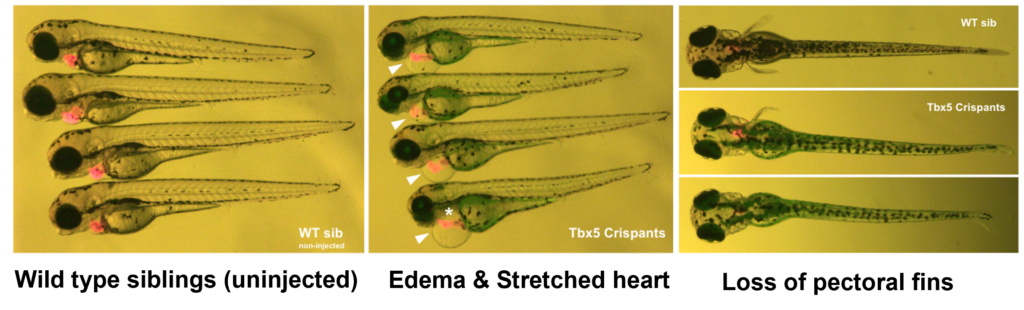
Fig. 1: See Crispant in action.
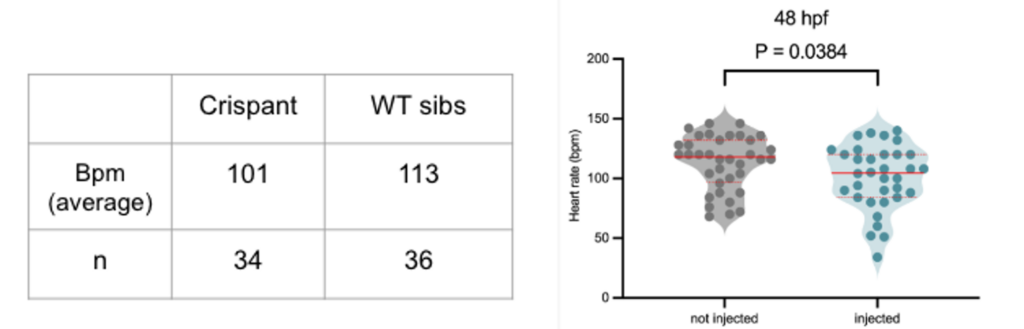
Fig. 2: A comparison of Heartstrings Mutant (tbx5). Witness how Crispant emulates both the stable KO and morphant, showcasing a slowed heartbeat, elongated heart structure, and the loss of pectoral fins.
Ready to Elevate Your Research?
Take the first step towards a more efficient genetic model creation process. Reach out to us with your inquiries, and our team will connect with you promptly.
FAQs About Crispants:
What are Crispants?
Crispants (CRISPR-Cas9 engineered animals) are genetically engineered animals with a single gene removed (or modified), which allows for the study of the effects of that gene on growth and development. Crispants have been used to explore the role of genes in diseases, such as cancer, and to test new treatments and drugs. They have also been used to develop animal models of human diseases, such as diabetes and heart disease.
- How do Crispants influence mutant alleles in the approach for zebrafish studies?
Crispants can be used to change the expression of a gene in zebrafish by targeting and deleting a specific section of the gene. By deleting this section, the gene can no longer produce the gene product, which causes the mutant allele to be expressed. This allows researchers to study the effects of this mutation on growth and development and to better understand the roles of protein-coding genes.
- What are the primary benefits of using Crispants for studying gene functions and creating custom mutations in zebrafish?
Crispants allow for precise manipulation of genes, provide access to understanding gene function and facilitate the study of differentially expressed genes. They also provide an affordable method of creating custom mutations in zebrafish, allowing for more experiments to be conducted in the same time frame. Also, Crispants can be used to create transgenic lines with the exact mutations necessary to explore certain diseases and treatments. - How does the InVivo’s proprietary Crispant approach differentiate from traditional CRISPR techniques in fast-tracking KO knock-in lines?
While traditional CRISPR methods might require almost a year, InVivo’s unique Crispant approach leverages deep learning methods to produce fast-track KO knock-in lines in less than a month. This enables researchers to swiftly generate transient knock-out models with high vivo editing efficiencies, which can then be used to establish long-term stable zebrafish knock-out models with specific genomic mutations.
- Can Scientists use Crispants to create mice with mutations that mimic those found in humans with diseases such as Alzheimer’s or Parkinson’s?
Yes, Crispants can be used to create mouse models with frameshift mutations that mimic those found in humans with diseases such as Alzheimer’s or Parkinson’s. In fact, by leveraging the zebrafish community and the insights from approaches in zebrafish research, Crispants have already been used to create mouse models with mutations that mimic those found in humans with these diseases.
References:
- https://pubmed.ncbi.nlm.nih.gov/30912049/
- https://www.ncbi.nlm.nih.gov/pmc/articles
- https://www.zeclinics.com/services-and-solutions/fast-track-knock-out-crispants/
