Advanced Drug Discovery & Development Services
More Preclinical CRO Services
Target-based vs. Phenotype-based
Drug Discovery
The drug discovery process for researching and developing new medicines is growing in difficulty and length. On average, it takes at least ten years and with an estimated average cost of billions of dollars to bring a new medicine to market [1,2]. The human and financial cost of failures is enormous due to the fact that thousands and sometimes millions of compounds may be screened and assessed early in the R&D process, but only a few will ultimately receive approval. For early-stage biotech and pharma companies, the return is even lower because they don’t have many compounds to screen.
A recent study of approved drugs revealed that Target-based Drug Discovery, or TDD, is underperforming relative to Phenotype-based Drug Discovery, or PDD. PDD focuses on ameliorating phenotypes to discover efficacious drugs, whereas TDD focuses on interactions with pre-identified drug targets. Looking broadly back over the past two decades, the PDD approach has resulted in twice as many approved new drugs as TDD[3]. Small and large pharma companies can benefit from broadening their PDD programs. By focusing on the phenotype, efficacious drugs are identified more quickly and readily, costing less time and money.
Target-based Drug Discovery
Phenotype-based Drug Discovery
Requires an identified target
No target identification needed
Unsophisticated cell lines /methods can be used
More complex genetic models are needed
Output is molecular and requires later efficacy testing
Output is functional, and may need later MMOA identification
Our Offerings
Two of our most common services are:
- compound testing (toxicity assessment and measuring compound effects);
- understanding biological functions of a disease gene.

Compound Efficacy Assessment
Testing new compounds and formulations to understand whether they are functionally effective on disease phenotypes.
- Dose Determination: Determine test dosage through evaluations of toxicity and solubility.
- Create assay systems and models: Genetically reproduce disease state or condition in the most well-suited animal model.
- Measure compound effects: Use a functional phenotype to generate data about compound efficacy.
- Data modeling: Sophisticated data analysis reveals whether candidate compounds are effective

Mechanism of Action Studies
Creating “Clinical Avatars” by inserting the disease gene variant into a model to create humanized animal model(s) that express human disease genes.
- Generate model(s): creating genetic models using CRISPR or other genome editing methods. Multiple genetic variants can be modeled to match population demographics.
- Perform genetic perturbations: RNAi treatments, protein degradation, and complementation studies are used to provide gene expression and genetic modifier screens
- Phenotypic profiling: Data generation and collection using optimized relevant assays (e.g., behavioral, metabolic) and transcriptional analysis.
- Understand the mechanism of actions: Analyzed data delivers information about molecular targets, biochemical pathways, and other MoAs.
Other Services and Testing
We also perform other specialized testing and analysis including:
- In Vivo Toxicity Testing: Conduct toxicity screening using zebrafish or C. elegans.
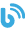
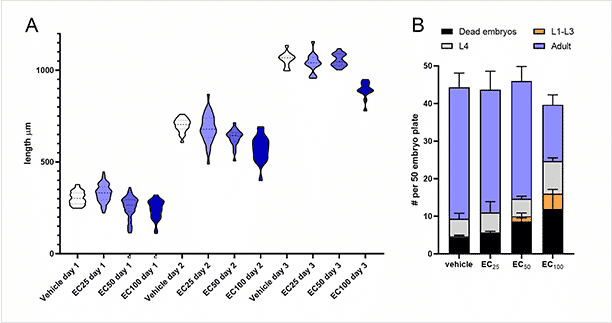
Data example: By combining a highly sensitive larval growth assay with a highly selective egg viability assay, we can refine the pool of possible compounds or dosages to those least (or most) likely to be toxic.
A) Larval growth assay violin plot shows distribution of worm length measured by high resolution imaging and automated worm detection. Darker blue indicates higher drug concentration. Day 1 = L2/L3, day 2 = L4, day 3 = adult.
B) Egg viability assay showing the average proportion outcomes for 50 embryos 3 days after laying. EC = effective concentration relative to maximal response of a fluorescent stress reporter. Learn more about our In Vivo Toxicity Testing.
- Lifespan and Healthspan Analysis: Conduct toxicity screening using zebrafish or C. elegans. Determine whether a compound can extend lifespan, promote a longer healthspan, and test the outcome on signaling pathways and cellular mechanisms promoting longer life using C. elegans
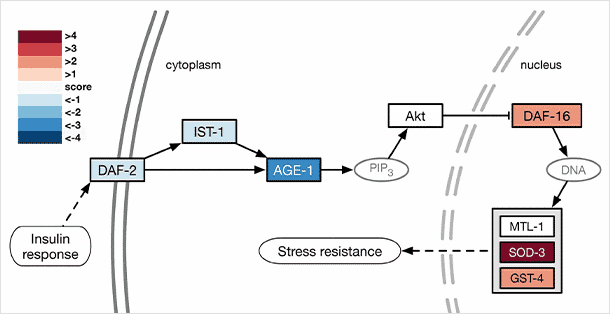
Mapping of gene expression data to longevity pathways in the KEGG database
Pathway shown represents changes of expression that might be observed under conditions of dietary restriction. Pathway components are color coded based on the level of up or down regulation of treated samples versus control.
Learn more about our Lifespan and Healthspan Analysis.
- Drug repurposing for rare genetic diseases: Screen market-available drugs and/or compound libraries to reveal the therapeutic best fit for rare genetic diseases.Contact Us:
References
- Biopharmaceutical Research & Development: The Process Behind New Medicines. PhRMA. 2015.
- Estimated Research and Development Investment Needed to Bring a New Medicine to Market, 2009-2018. JAMA. 2020;323(9):844-853. doi:10.1001/jama.2020.1166
- How Were New Medicines Discovered?Swinney, David C., and Jason Anthony. 2011. Nature Reviews Drug Discovery.
Ready to get started?
Ready to connect with us to learn more about working with our company or our technology?
Submit your inquiry below & we will get back to you soon.