Humanized Models For Disease Research And Treatment Insights
More Disease Modeling Services
Genome Edited Animal Models For Studying Your Gene Of Interest
“When people talk about ‘model organisms,’ many people think that the animal has to ‘model’ the human disease condition to have any impact. We argue that this is not the case. Simply studying what the homologous gene does in the context of the humanized zebrafish model organism can provide important insights into disease pathology.
Humanized animal models, including humanized models for disease research and treatment insights, play a crucial role in understanding genetic mutations and disease biology discovery. “Rare genetic variants can be modeled in a number of different model organisms, informing causality between a particular variant and a pathogenic phenotype and offering clinicians and their patients hope for a diagnosis, insight into the mechanisms and pathways involved in a rare disease, and ways it might be treated.” Neff, E.P. Model matchmaking. Lab Anim 50, 39–42 (2021).

Using CRISPR and other genome editing methods, InVivo Biosystems can humanize or genetically engineer zebrafish and C. elegans into powerful humanized animal models for better understanding genetic mutations and disease biology discovery.
- HElio Ortholog Gene Finder: Find a usable model for studying your gene of interest.
- Disease Gene Finder: Linking human disease genes and C. elegans genes and their corresponding phenotypes.
- Publications for disease research using C. elegans. Top disease research areas using C. elegans model.
Methods For Humanization
There are two types of humanizing changes that can be made:
- Whole gene humanization – swapping in human sequence for animal gene
- Point mutation humanization – inserting clinical variation into animal’s gene
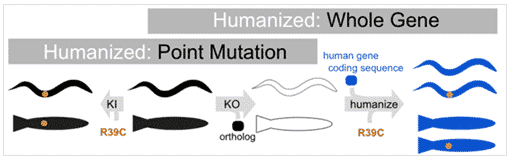
Humanization with Point Mutation
Humanization with Whole Gene
Speed
++
+
Cost
$
$$
Accuracy
+
+++
Drug Screening
+
+++
Whole Gene Humanization
Whole gene humanization is often used to model the effects of the clinical variant.
- The human coding sequence is used to replace the animal’s coding sequence.
- CRISPR techniques are use to remove the entire coding sequence of the animals version of the gene.
- A plasmid with homology flanked cargo is used to bring in the human coding sequence.
We know we are looking at conserved biology between animals and humans when the human gene rescues the function of the gene-deleted KO animal. Next we put clinical variations into the humanized locus which allows us to examine any variation (rare or common) for its functional contribution to normal biology. When we observe abnormal biology in a clinical variant, the system becomes a Human Avatar to screen drugs and find therapeutics that can restore normal function.
We have found whole gene humanization can create a platform for highly translatable results in a model organism. The gene humanization method we use replaces the ortholog locus with human cDNA. The cDNA is sequence-optimized for expression in the animal.
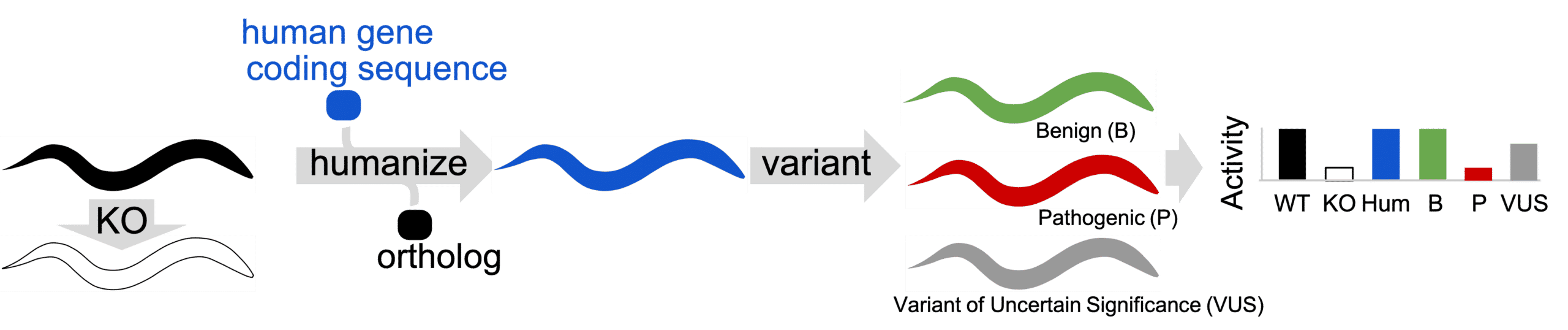
Gene-swap humanization technique. Ortholog gene is removed from animal (KO) to determine if a defective phenotype is present. Ortholog is replaced with human coding sequence to create a gene-humanized animal. Variants are installed in gene-humanized locus to create Patient Avatars. Humanized animals in Patient Avatar format are phenotyped for activity defects.
Point Mutation Humanization
Point mutation humanization is a simpler, quicker and fairly effective – CRISPR techniques to replace a single amino acid with the suspect amino acid seen in the patient. The clinical variant seen in the patient is installed in the animal’s version of the gene.
Note, that this only works if sequence conservation is high and even then some variants that are pathogenic may not be at conserved positions.
Modeling STXBP1 Patient Variants in C. elegans
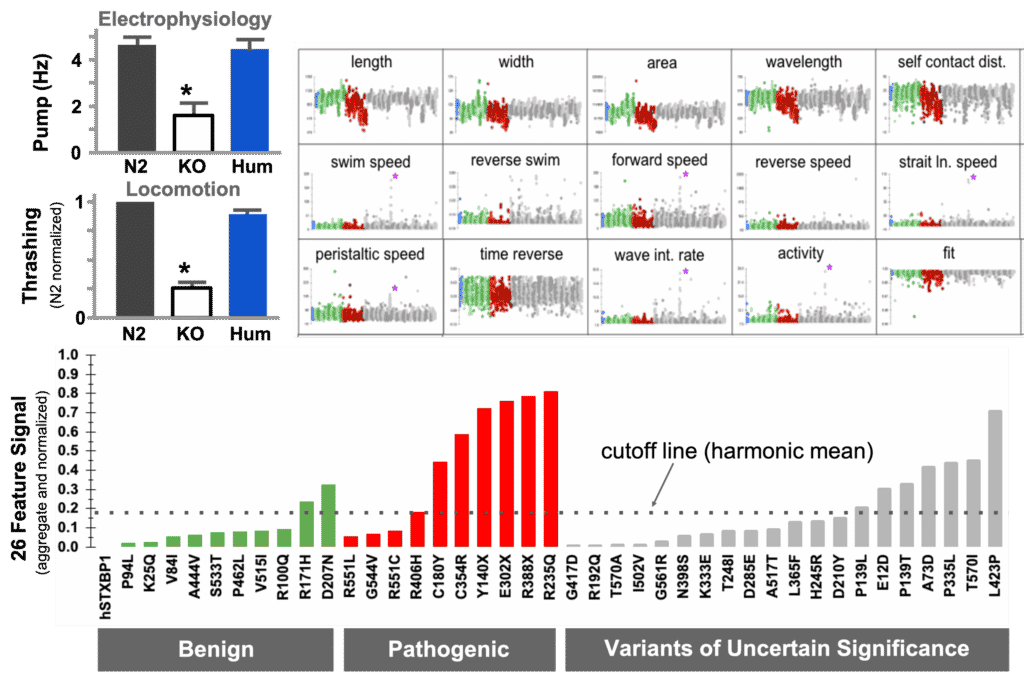
In this example, the human coding sequence of STXBP1 was able to restore function when inserted as whole-gene humanization of the unc-18 locus. Referenced against a set of known pathogenic and benign variants, we were able to detect pathogenicity in a set of VUS.
The diagnostic curve for VUS assessment. (a) Transgene rescue demonstration by electrophysiology. (b) Transgene rescue demonstration by movement in liquid. (c) Deep phenotyping parameters for movement on solid surface compare benign (green), pathogenic (red) and VUS (grey). d) diagnostic curve for pathogenicity assessment of VUS where values above harmonic mean indicate possible pathogenicity in the strain.
The video below shows that the human coding sequence of STXBP1 is inserted into the native locus of a C. elegans ortholog gene. The worms version of the gene is replaced with a human coding sequence.
– Left: the gene-swapped STXBP1 sequence functions and shows a significant level of activity.
– Middle: In contrast, the knock-out shows very little activity.
– Right: for the R406H genomic variant, its activity is somewhere in between the “humanized” and “knock-out” strain.
96 point mutations in 1 gene. All 96 mutations were created in the STXBP1 gene (which is associated with epilepsy in humans) via CRISPR. The worm homolog of STXBP1, unc-18, causes incoordination and a near-complete lack of pharyngeal pumping when knocked out. The functionality is restored by replacing the worm gene with the coding sequence for human STXBP1.
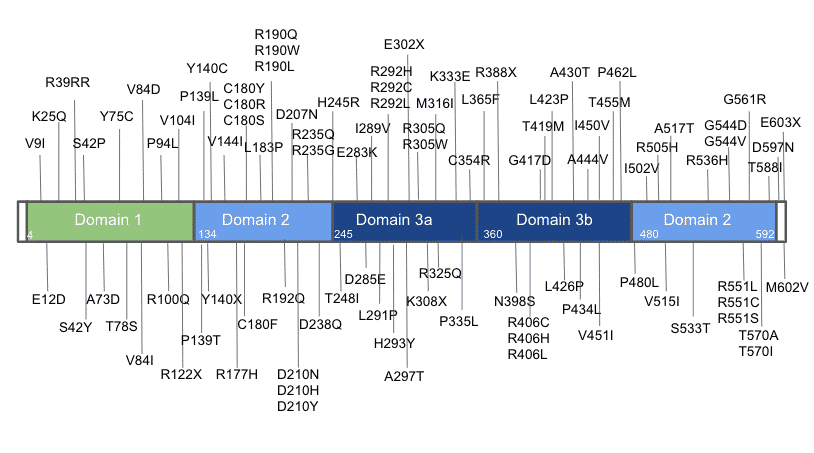
Modeling STXBP1 Patient Variants in Zebrafish
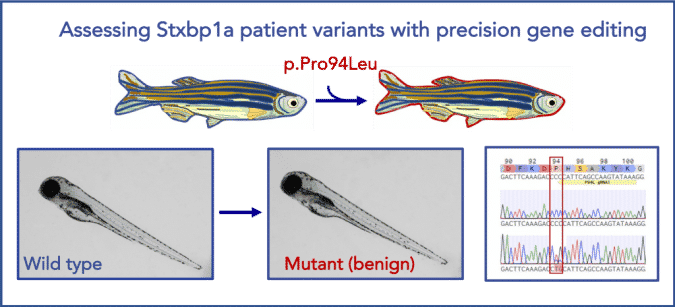
Illustration of a successful precise point mutation of stxbp1a in zebrafish. stxpb1a is a highly conserved zebrafish ortholog of human STXBP1 (87% identity). Using CRISPR/Cas9 technology, we were able to precisely generate a benign patient mutation at the conserved amino acid residue (CCC>CTG, p.P94L).
Who We Serve
We work with scientists seeking new or alternative ways to approach their research.
- R&D researchers who need to test their hypotheses before moving to a mammalian system.
- Pharmaceutical companies who need to quickly understand a compound’s mechanism of action as well as cytotoxic effect, CC50, IC50, SI, and etc.
- Animal-conscious organizations who need to perform proof-of-concept validation.
- Researchers studying longevity and senescence who want to eliminate the maintaining and handling of mouse model for months.
Resources
InVivo Biosystems is ready to help you with your COVID-19 research! We offer our genetic expertise in the engineering of humanizations, fluorescence markers, luciferase, activity reporters, and conditional expression to help you interrogate drug and vaccine activity, binding interactions, mechanism of action, and more.
Humanized Models For Disease Research and Treatment Insights
1. What are Humanized Models in Disease Research?
Humanized models are innovative tools in biomedical research that involve modifying animal models, such as mice or C. elegans, to carry human genes, human cells, or human immune system components. These models are pivotal for studying complex human responses to diseases and treatments in a controlled environment. By incorporating human donor genetic material or humanized immune system mice (e.g., HIS-NSG mice), researchers can observe the effects of diseases and treatments with greater relevance to human physiology, leading to more accurate predictions of clinical outcomes in drug discovery and gene therapy.
2. How do Humanized Models Improve Drug Discovery Processes?
Humanized models significantly enhance the drug discovery process by providing a more accurate representation of how therapeutic agents interact with human biology. These models, including humanized mouse models and patient-derived xenografts (PDX), allow for the evaluation of drug efficacy and toxicity in a system that closely mimics human disease states. Utilizing vivo models and preclinical models enables the identification of potential adverse reactions and cytokine release syndrome risks before clinical trials, optimizing drug screening protocols and improving the safety and effectiveness of new treatments.
3. What Role Do Humanized Models Play in Understanding Infectious and Autoimmune Diseases?
Humanized models play a crucial role in understanding infectious diseases and autoimmune diseases by simulating the human immune response within an animal model. Through the introduction of human immune cells or immune system components into immunodeficient mouse models, researchers can study the host immune interactions, immune cell chimerism, and the development of diseases within a human-like immune environment. These insights are essential for developing targeted therapies that can modulate the immune response, offering hope for patients with conditions previously considered untreatable.
4. How are Humanized Models Developed for Disease Research?
The development of humanized models for disease research involves several sophisticated techniques, including CRISPR/Cas9 for genome editing and the engraftment of human cells or tissues into immunodeficient mice. Whole gene humanization swaps animal genes for their human counterparts, while point mutation humanization introduces specific clinical variants into the animal genome. This precise genetic engineering, exemplified by the creation of humanized mouse models like HIL-NSG mice and humanized immune system mice, allows for the detailed study of human gene function, immune cell subsets, and disease mechanisms.
5. What are the Challenges and Limitations of Using Humanized Models in Research?
Despite their significant advantages, humanized models also face challenges and limitations. The complexity of the human immune system and the intricate interplay of immune cell subsets may not be fully replicated in animal models. Long-term studies and weeks post-engraftment observations are required to understand the immune cell composition and myeloid cell reconstitution dynamics. Moreover, ethical considerations and the need for human donor tissues can limit the availability of certain models. Researchers must also navigate the genetic modifications and natural killer cell activities that can differ between the model and human conditions, potentially affecting the accuracy of study results.
Ready to get started?
Ready to connect with us to learn more about working with our company or our technology?
Submit your inquiry below & we will get back to you soon.




