Here at InVivo Biosystems we spend a lot of time thinking, and talking, about genome editing. We thought it would be fun to compile a list of our frequently used terms – so here they are, our ABCs of genetic research.
A: Animal Models
Animal models are organisms (such as mice, rats, fruit flies, worms, zebrafish, frogs and apes) that are used in biomedical research to mimic humans. There are four types of commonly used model organisms: disease induction models, xenograft models, inbred strains, and transgenic models. These models enable researchers to gain insight into biological processes such as development as well as disease mechanisms.
B: Base Pair
Base pairs are the two chemicals that are bonded together, often referred to as one of the “rungs on the DNA ladder”.
C CRISPR
CRISPR is an acronym for clustered regularly interspaced short palindromic repeats – a family of DNA sequences that are the ‘hallmark’ of a bacterial defense system. These repeating sequences of genetic code have served as the basis for CRISPR-Cas9, a technology which enables researchers to edit genes.
D: DNA
Short for Deoxyribonucleic acid, DNA is the hereditary material in humans and nearly all other organisms. DNA takes the shape of a double-helix thanks to its two polynucleotide chains which coil around each other.
E: Exome
Exome is the collective of all ‘exons’ in one’s genome. Exons are the sequences which code for proteins – they determine an organism’s phenotype.
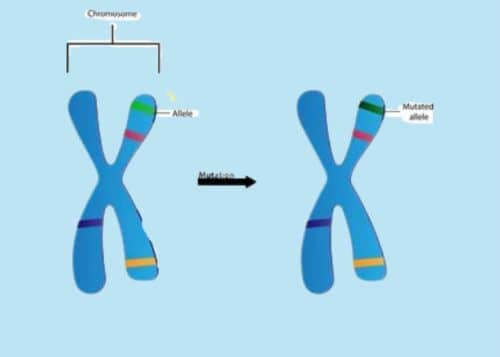
Figure 1. Gene Allele Illustration.
F: Floxed allele
Floxing is the sandwiching of a DNA sequence between two lox P sites. In animal models, floxed alleles enable researchers to make knockout models (deletion of genes in site and time-specific manner).
G: Genotyping
Genotyping is the process of determining an individual’s genetic sequence variants.
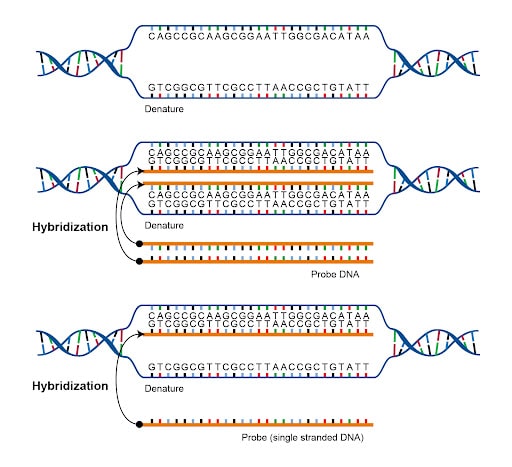
Figure 2. Illustration of hybridization.
H: Hybridization
Hybridization refers to the process in which two single-strand DNA or RNA molecules combine through base pairing, resulting in a double-stranded molecule. This process can also be reversed. As such, this is part of many lab techniques related to specific DNA samples.
I: In vivo
Latin for ‘within the living’, in research, in vivo refers to studies that occur within a living organism. Since they occur within the organism, in vivo studies enable researchers to better understand disease pathology. Learn more about the ins (vivo, vitro, silico) of research.
J: Junk DNA
Junk DNA is the region of DNA that does not encode proteins (this makes up nearly 98% of our DNA).
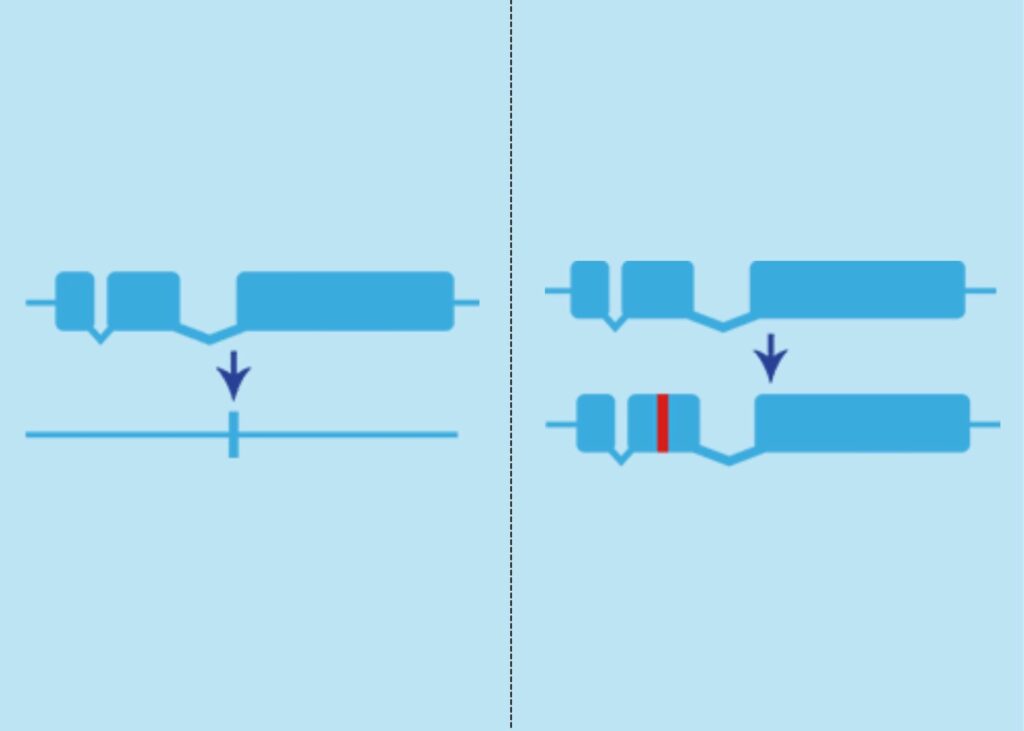
Figure 3. Knockout (left) vs. Knockin (right).
K: Knockout (KO)/ Knock-in (KI)
KO and KI technologies are used to genetically alter model organisms. This process involves deactivating or removing a gene (KO), or inserting a new gene (KI).
L: Locus
The specific location of a gene or DNA sequence on a chromosome.
M: MosSCI
MosSCI is a method of inserting transgenes into a model organisms’ genome at the designated locus. This method works by exploiting the cell’s natural repair system – creating a double-strand break at a defined site and utilizing an exogenous DNA template which the cell will incorporate as it repairs.

Figure 4. C. elegans
N: Nematode (our favorite is the C. elegans!)
Nematode is the latin word for worm, and while there are many varieties used in research our personal favorite is the C. elegans. C. elegans (Caenorhabditis elegans) are one of the most commonly used model organisms because their simplicity enables researchers to gain invaluable insight into how genes affect development and behavior.
O: Ortholog
Orthologs are genes that have evolved from a common ancestor. Generally, these genes retain the same function, which enables researchers to conduct studies on model organisms‘ genes and translate the results to humans.
P: Phenotype
A phenotype is an observable trait (eye color, height, etc.) that can be determined by an individual’s environment, or genotype (genetics).
Q: Quantitative PCR (qPCR)
A laboratory technique that is used to characterize and quantify nucleic acids in real-time.
R: Replication
DNA replication is the process of a double-stranded DNA molecule producing two identical replications of itself.
S: Site-directed mutagenesis
A method to make specific DNA alterations (insertions, deletions, and substitutions) in double-stranded plasmid DNA.
T: Transgenics
An organism that has had genetic material artificially introduced through techniques such as CRISPR.
U: Upregulation
The upregulation of gene expression is the process in which the cell increases the production of genetic material (protein or RNA). This process is used to adapt to new environmental stimuli and trigger developmental pathways.
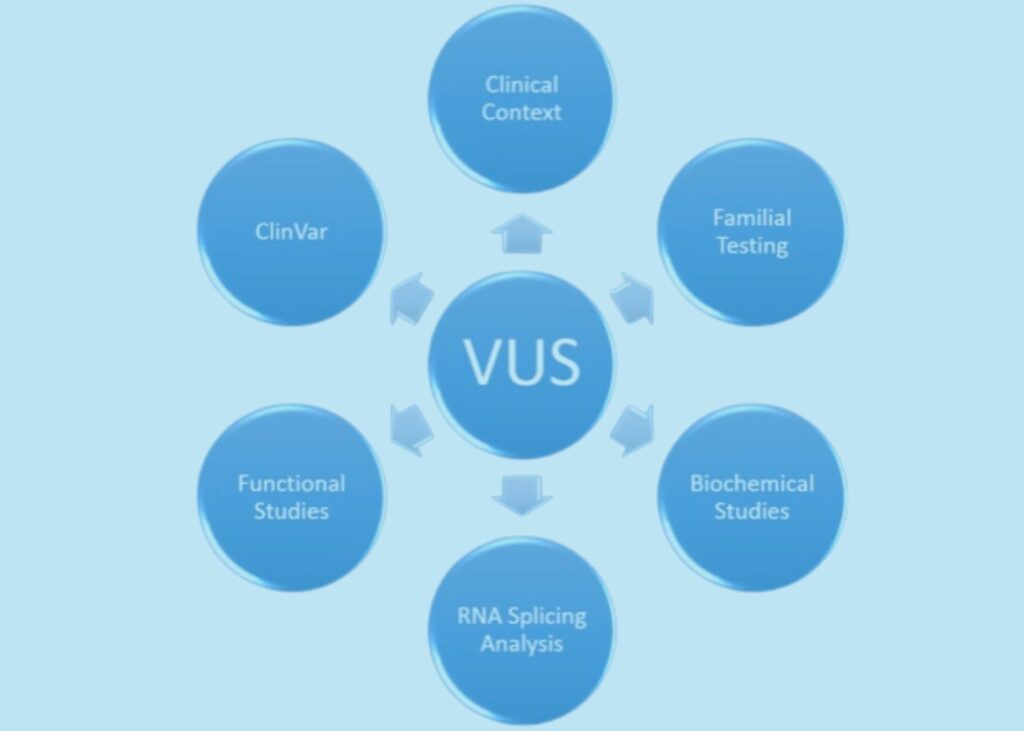
Figure 5. Ways of evaluating variants of unknown significance (VUS).
V: Variants (VUS)
A genetic variant (change) whose impact on an individual’s health and/or function is unknown.
W: Wild type
An organism that has not undergone genetic manipulation, or shows genetic variation, but rather is ‘normal’ for their species.
X: Xenograft
A xenograft is the implantation of an organ, tissues, or cells from one species into another – these are particularly powerful for modeling cancers where a patient’s tumor can be transplanted into an animal model.
Y: Y-chromosome
The Y chromosome is one of the two sex chromosomes in humans (the other being the X-chromosome). whether a person has an XX (biologically female) or XY (biologically male) pairing can be a risk factor for certain genetic diseases.

Figure 6. zebrafish.
Z: Zebrafish
The zebrafish (Danio rerio) is a popular model organism. A freshwater fish, its small size, low maintenance, and high translatability with humans make them a go-to choice for many researchers (particularly for toxicity and drug screening).
Looking for a precise and reliable way to introduce specific genetic modifications into your cells? Our CRISPR Gene Knockin service may be just what you need! Our experienced team uses cutting-edge technology to precisely insert genetic material into your cells, allowing you to study the effects of specific gene modifications. Whether you’re working on basic research or drug discovery, our CRISPR Gene Knockin service can help you achieve your research goals. Contact us today to learn more about our services!



