Summary
Dr. Robyn Tanguay is a distinguished professor at Oregon State University, and a collaborator with InVivio Biosystems. Dr. Tangauy utilizes zebrafish to study toxicology, and last month received a $7 million grant from the National Institute of Environmental Health Sciences to study the biological impacts of chemicals. In a recent publication, Dr. Tanguay worked with Dr. Maret Traber to create a new fluorescent-tagged zebrafish line to study the effects of Vitamin E on embryonic development. This article will give an overview of the Traber Lab’s work, and discuss: ‘why zebrafish’?
The Traber Lab has done extensive research on the effect of Vitamin E, and has shown that its deficiency causes deregulation in energy metabolism, gene expression networks, and the nervous system during development, as well as increased morphologic abnormalities and mortality (McDougall et al., 2017; Head et al., 2021b; Miller et al., 2021; Head et al., 2020). In a recent publication the Traber Lab, in collaboration with InVivo Biosystems, utilized CRISPR-Cas9 technology, and the red fluorophore mScarlet, to generate a Ttpa-tagged zebrafish model to study Vitamin E in early embryonic development.
Glossary:
- Alpha-tocopherol transfer protein = the vitamin E regulatory protein
- TTPA = a-tocopherol protein in humans, mice, and rats
- Ttpa = a-tocopherol protein in zebrafish
Vitamin E deficiency in mothers and babies can lead to low birth weight and an increased risk of miscarriage (Evans & Bishop, 1922; Shah et al., 1987). This deficiency can be caused by genetic defects in the Vitamin E regulatory protein, a-tocopherol (referred to as TTPA in humans and Ttpa in zebrafish). TTPA is primarily located in hepatocytes (liver cell), and human yolk sac and placental lining (Head et al., 2021a). But, despite the established link between Vitamin E and embryonic development, study of TTPA in the developing embryo is limited. To address this unmet need, Dr. Tanguay and the Traber lab injected zebrafish embryos with the coding sequence for the fluorescent mScarlet along with a cas9 protein, raised them to adulthood, and then breed them to establish a new zebrafish line. The result is a model which expresses mScarlet fluorescence in the presence of Ttpa, allowing researchers to easily track VitE and Ttpa [Figure 1].
Figure 1. In vivo fluorescence detected in 7 and 14 days post-fertilization RedEfish. (Head et al., 2021a). (A) mScarlet expression was unique to the olfactory pits (op) found in the RedEfish; however, some autofluorescence was detected in the pigment cells surrounding the eyes (e) in all embryos. (B) mScarlet expression in the digestive tract, specifically the liver (*). (C) More punctate expression was observed in the anterior gut region. (D) Punctate expression was observed also in the tail fin (*). (E) In late stage development (14dpf) mScarlet expression was identified in the caudal vertebrae and dorsal root ganglia (drg) and the caudal vein plexus (cpv). 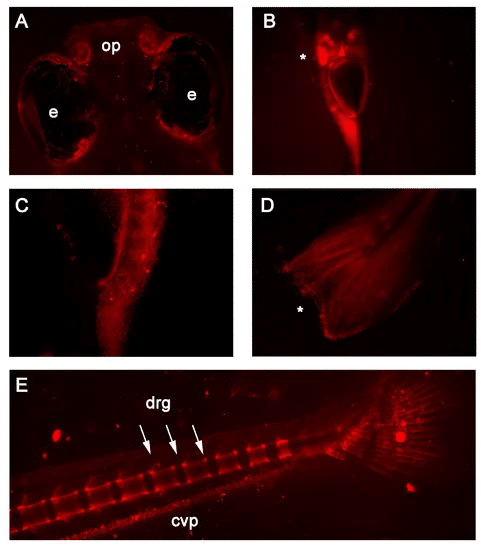
The Traber Lab chose to use zebrafish as their model organisms because zebrafish are a highly advantageous model for studying embryos, thanks to their transparency and external development, as well as their ease to maintain [Figure 2]. Zebrafish are also an excellent model for this type of research because they are so established, having been used in numerous birth defect studies (Head et al., 2021a). Lastly, zebrafish are easy to genetically manipulate, which means this new line, (called the RedEfish mScarlet:GSG-T2A:Ttp Transgenic line) will enable other research groups to carry-out the much needed studies on Ttpa localization, and VitE trafficking.

Figure 2. A few key points of why zebrafish are excellent models for Vitamin E regulation.
References
- Evans, H. M., & Bishop, K. S. (1922). ON THE EXISTENCE OF A HITHERTO UNRECOGNIZED DIETARY FACTOR ESSENTIAL FOR REPRODUCTION. Science (New York, N.Y.), 56(1458), 650-651. https://doi.org/10.1126/science.56.1458.650
- Head B, La Du J, Barton C, Zhang J, Wong C, Ho E, Tanguay RL, Traber MG. (2021a) RedEfish: Generation of the Polycistronic mScarlet: GSG-T2A: Ttpa Zebrafish Line. Antioxidants; 10(6):965. https://doi.org/10.3390/antiox10060965
- Head, B., Ramsey, S. A., Kioussi, C., Tanguay, R. L., & Traber, M. G. (2021b). Vitamin E Deficiency Disrupts Gene Expression Networks during Zebrafish Development. Nutrients, 13(2), 468. https://doi.org/10.3390/nu13020468
- Head, B., La Du, J., Tanguay, R.L. et al. Vitamin E is necessary for zebrafish nervous system development. Sci Rep 10, 15028 (2020). https://doi.org/10.1038/s41598-020-71760-x
- McDougall, M., Choi, J., Kim, H. K., Bobe, G., Stevens, J. F., Cadenas, E., Tanguay, R., & Traber, M. G. (2017). Lethal dysregulation of energy metabolism during embryonic vitamin E deficiency. Free radical biology & medicine, 104, 324-332. https://doi.org/10.1016/j.freeradbiomed.2017.01.020
- Miller, G. W., Labut, E. M., Lebold, K. M., Floeter, A., Tanguay, R. L., & Traber, M. G. (2012). Zebrafish (Danio rerio) fed vitamin E-deficient diets produce embryos with increased morphologic abnormalities and mortality. The Journal of nutritional biochemistry, 23(5), 478-486. https://doi.org/10.1016/j.jnutbio.2011.02.002
- Shah, R. S., Rajalakshmi, R., Bhatt, R. V., Hazra, M. N., Patel, B. C., Swamy, N. B., & Patel, T. V. (1987). Vitamin E status of the newborn in relation to gestational age, birth weight and maternal vitamin E status. The British journal of nutrition, 58(2), 191-198. https://doi.org/10.1079/bjn19870086



