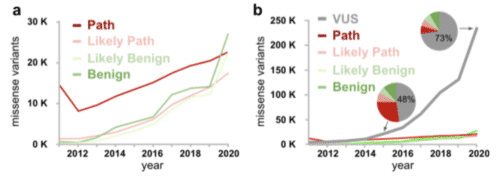
We have found whole gene humanization can create a platform for highly translatable results in a model organism. Currently there is a big need for functional studies of Variants of Uncertain Significance (VUS).
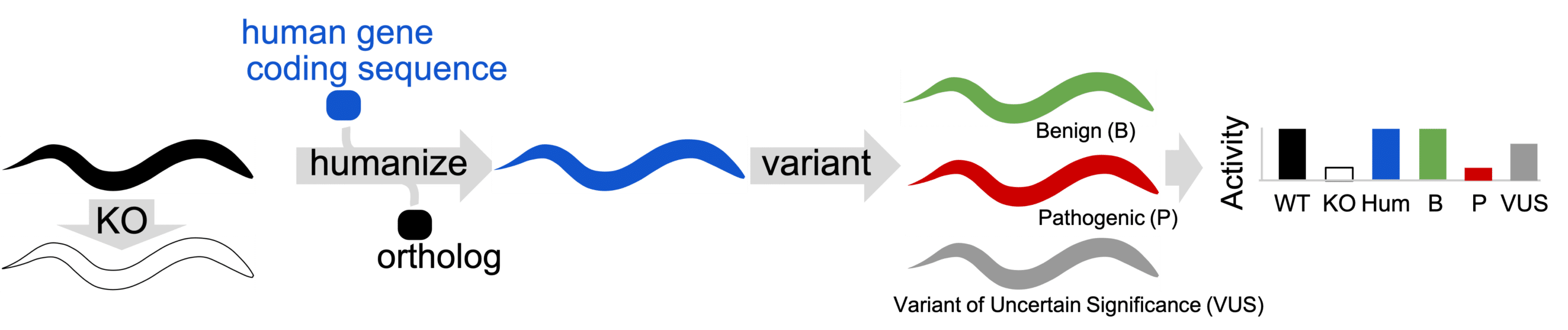
To address this need of the clinic, we can utilize functional studies in small animal models. To increase the relevance of the data into translatable results, we can use gene humanization. The gene humanization method we use replaces the ortholog locus with human cDNA. The cDNA is sequence-optimized for expression in the animal.
Two conditions must be satisfied for the system to be successful.
- Knock-out must have a phenotypic consequence. This may be an overt behavioral phenotype (eg. locomotion) or a clear expression response (decreased or increased gene expression).
- Human cDNA must restore normal function. The recoded sequence of the human gene inserted as coding sequence replacement must lead to restoration of normal function.
When these conditions are met, the system can be used to understand if pathogenicity is occurring in a clinical variant.
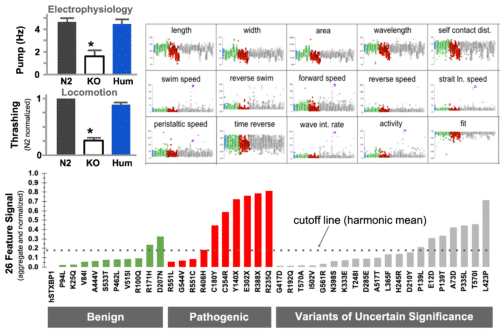
The STXBP1 gene (a leading cause epilepsy)
In an example with the STXBP1 gene, we have obtained satisfactory results. The human coding sequence of STXBP1 was able to restore function when inserted as gene replacement of the unc-18 locus. Referenced against a set of known pathogenic and benign variants, we were able to detect pathogenicity in a set of VUS.
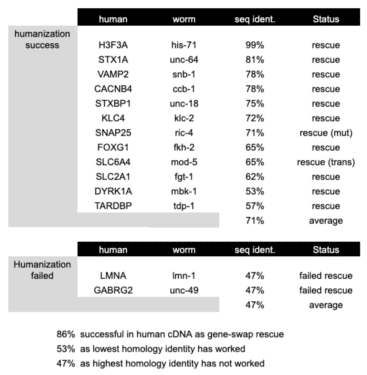
Generalizable Method (Determining the boundaries of success)
So far the results with this and other genes are quite encouraging. Genes with sequence similarity over 50% have not yet failed to create a functional assessment platform. When we look at the success rate for 15 completed projects, we can see that more than 85% of the genes we have tried are yielding a humanized system that can rescue a phenotypic defect of the null.
In future work, once humanized animal models are found to exhibit pathogenic behavior, drugs can be explored for capacity to restore phenotypic activity back to wild type. As a result, we are eager to find collaborators/partners who can help us in the endeavor to start helping with patient diagnostics and drug discovery in Rare Disease.