Summary:
Gut microbiota impacts nearly every aspect of health, but researching it can be tricky due to differences between humans and commonly used model organisms, as well as interindividual variability in humans. A relatively new model, zebrafish have become a gold-standard model for biomedical research, and are especially used to study host-microbe interactions. In this article we will discuss zebrafish’s contribution to recent microbiota research and the features that make them particularly well-suited for studying the host-microbe interactions.
As we discussed in the first part of this blog series, zebrafish offer many benefits as an alternative model in many studies: their small size and easy maintenance makes them cost effective, and significant homology to other vertebrates means that zebrafish research is easily translated to humans. One area where zebrafish can be particularly beneficial is the study of host-microbe interactions, as they provide a fast, straightforward way to examine the role of resident microbes during development.
Host-microbe interactions have been studied for as long as we knew that microbes existed, though until relatively recently this work focused on pathogenic interactions and ignored the beneficial effects of host-associated microbes. Much of the early development of zebrafish as a model for host-microbe interactions was led by Karen Guillemin and her laboratory at the University of Oregon and John Rawls and his laboratory at Duke University in the early 2000s. Since then, other laboratories across the world have used zebrafish to study host-microbe interactions.
Why Study Host-Microbe Interactions?
Interactions between microbes and their host are essential to the health and longevity of the host [Figure 1]. A person’s gut microbiota communication is in constant flux depending on age, diet, stress, environmental interactions such as travel, prevalence of pets, social interactions, and many other factors – at times, however, the change in gut microbiota is linked with diverse conditions, and this is known as dysbiosis. In addition to generating unpleasant intestinal symptoms like diarrhea, cramping, indigestion, bloating and constipation (Petra et al., 2015; Carding et al., 2015), dysbiosis is also a major risk factor for chronic illnesses such as IBS (Irritable Bowel Syndrome), Crohn’s disease, ulcerative colitis, and even neurological disorders including autism spectrum disorder and Parkinson’s disease (Parkin et al., 2021).
Recently, there is expanding awareness of how gut dysbiosis can affect the brain and peripheral nervous system both directly and indirectly via “the gut-brain axis.” It has been found that when a person is in a state of dysbiosis they report higher rates of brain fog and depression (Appleton, 2018).
The microbiota can also significantly impact the response to diverse pharmaceuticals. For example, one recent study showed that when 271 drugs were incubated with gut microbes, 176 of them metabolized to such an extent that the level of active drug dropped over 20% (Zimmermann et al., 2019). Similarly, interactions between drugs and the gut microbiota can transform some drugs into toxic metabolites – for instance, the most commonly reported adverse effect comes from non-steroidal anti-inflammatory drugs (NSAIDs) such as ibuprofen, which are known to cause gastroduodenal mucosal lesions in approximately 50% of regular users (Dikeocha et al., 2022). While NSAIDs mechanism are not fully understood, it is thought that this high development of lesions is due to NSAIDs modifying the growth and balance of microbe composition, causing changes in the relative abundance of the bacterial strains involved in drug absorption and metabolism (Maseda & Ricciotti, 2020). Therefore, studying host-microbe interactions is important for many responses, including understanding the mechanisms behind immune responses and longevity, which can enable the creation of safer, more effective drug therapeutics.
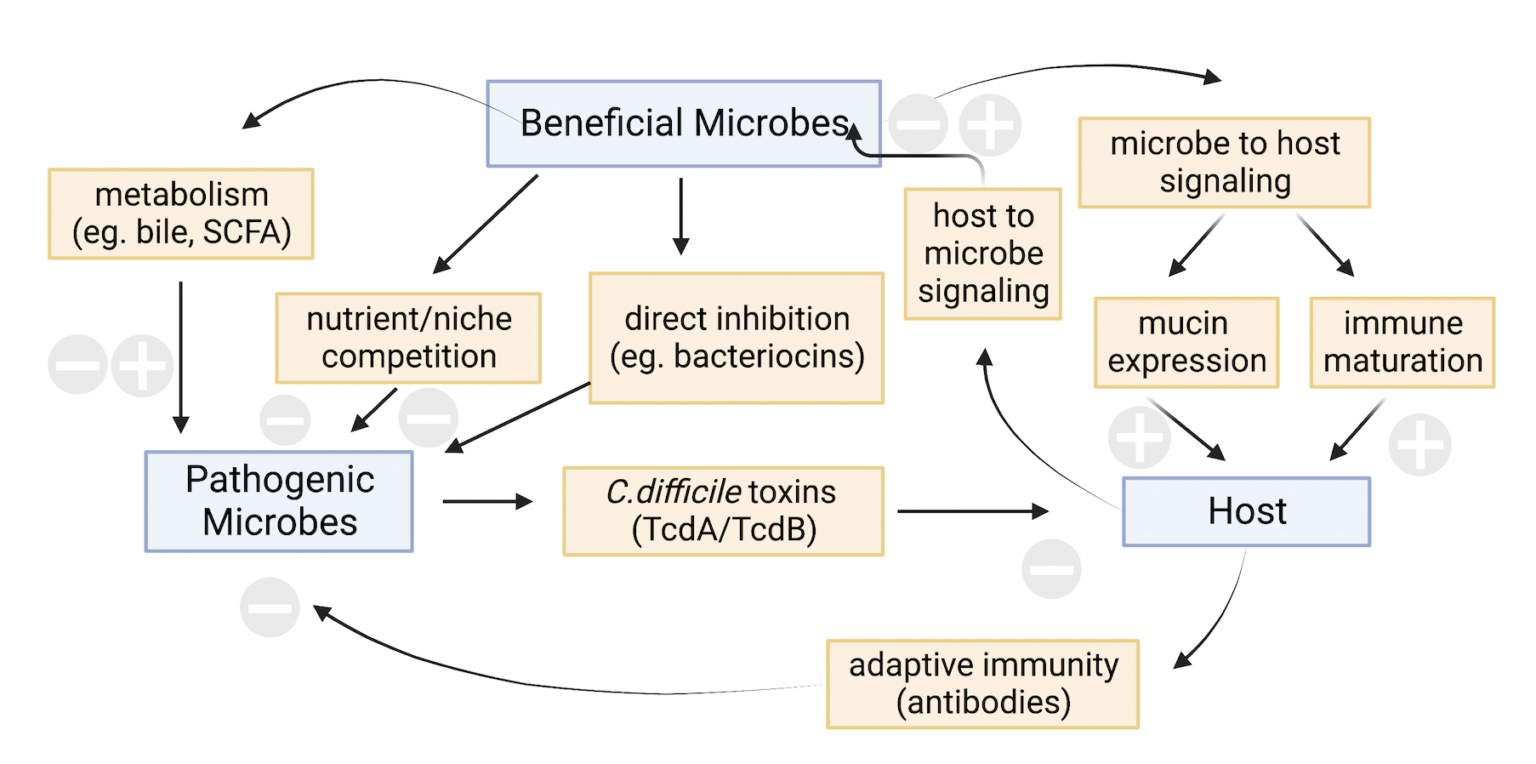
Figure 1. Representative interactions among beneficial microbes, pathogens, and the host (created in BioRender).
Zebrafish as Model for Host-Microbe Interactions
Advantages
Zebrafish has many notable benefits for this type of research. For example, it is relatively easy to raise germ-free zebrafish, or zebrafish with defined microbial communities, which is critical for understanding which microbes are required for host features like immune responses, gene expression, behavioral responses, and drug interactions (Tan et al., 2019).
Zebrafish also reach adulthood quickly (90 days), can reproduce every 10 days, and yield large broods (200-300 eggs per week). The ability of a single pair of parents to produce hundreds of genetically similar offspring makes them particularly valuable in microbiota research where controlling the genetic differences between individuals is critical for separating the influence of the microbiota from host genetic factors. Transparent zebrafish embryos and larvae make it easy to study the early embryo and early postnatal development, and these are critical periods for many host-microbe interactions, as both the host embryo and its associated microbial community are changing rapidly. The ability to conduct this early-life observation and manipulation of host-microbe interactions in zebrafish is particularly notable because the relative difficulty of experiments using mouse pups means that studies in mammals have largely focused on host-microbe interactions in adults. For example, studying the gut-brain axis in mammals can require complex dissection and other techniques; this is often not the case in the zebrafish, however, where gut-brain studies can be accomplished in an intact animal.
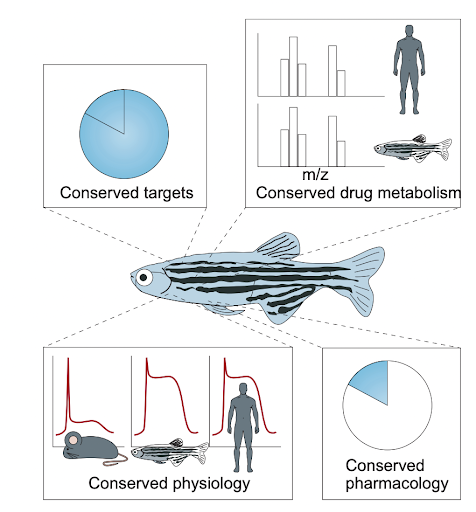
Figure 2. Zebrafish for studying human diseases (MacRae & Peterson, 2015).
Zebrafish are also vertebrates, and are similar to humans in critical ways: zebrafish have a highly-conserved immune system, lipid metabolism, nervous system, cell proliferation/apoptosis, carbohydrate metabolism, and glucose metabolism, among other systems, which means that many of the processes that occur in the human body can also be studied in zebrafish [Figure 2]. Though zebrafish may not be more similar to humans than rodents, zebrafish have a combination of similarities and experimental advantages that makes them particularly well-suited for certain experiments. For example, zebrafish are diurnal (active during the day and asleep at night). Therefore, studies in fish are easier to schedule and may be easier to translate to humans than those confounded by different circadian rhythms in nocturnal rodents.
Despite differences in some organ systems between zebrafish and mammals, such as in the gastrointestinal system, the functional units of these systems are highly conserved (Flores et al., 2020). For example,the intestinal bulb in zebrafish is similar to the mammalian small intestine and the rest of the gut is similar to the colon, analogies that are useful in host-microbe interaction studies (Flores et al., 2020). Unlike invertebrates, zebrafish also have both adaptive and innate arms of the immune system. Compared to mammals, adaptive immunity develops later in zebrafish, typically 4-6 weeks after hatching and microbial colonization. However, this difference makes zebrafish useful for separating pathways involving the two arms of immunity, which can be more difficult to do in mammals.
Zebrafish are also highly visual animals, while mice rely heavily on their sense of smell. This difference in visual vs. olfactory dominance means that zebrafish can be used to study complex behavioral patterns and high-throughput, complex behavioral analysis. Zebrafish exhibit many of the same types of basic behaviors as mammals, but they are easier to study with higher throughput (Levraud et al., 2022). For example, zebrafish can be utilized to study the effect of gut microbiota composition on social behavior, and fear/anxiety responses.
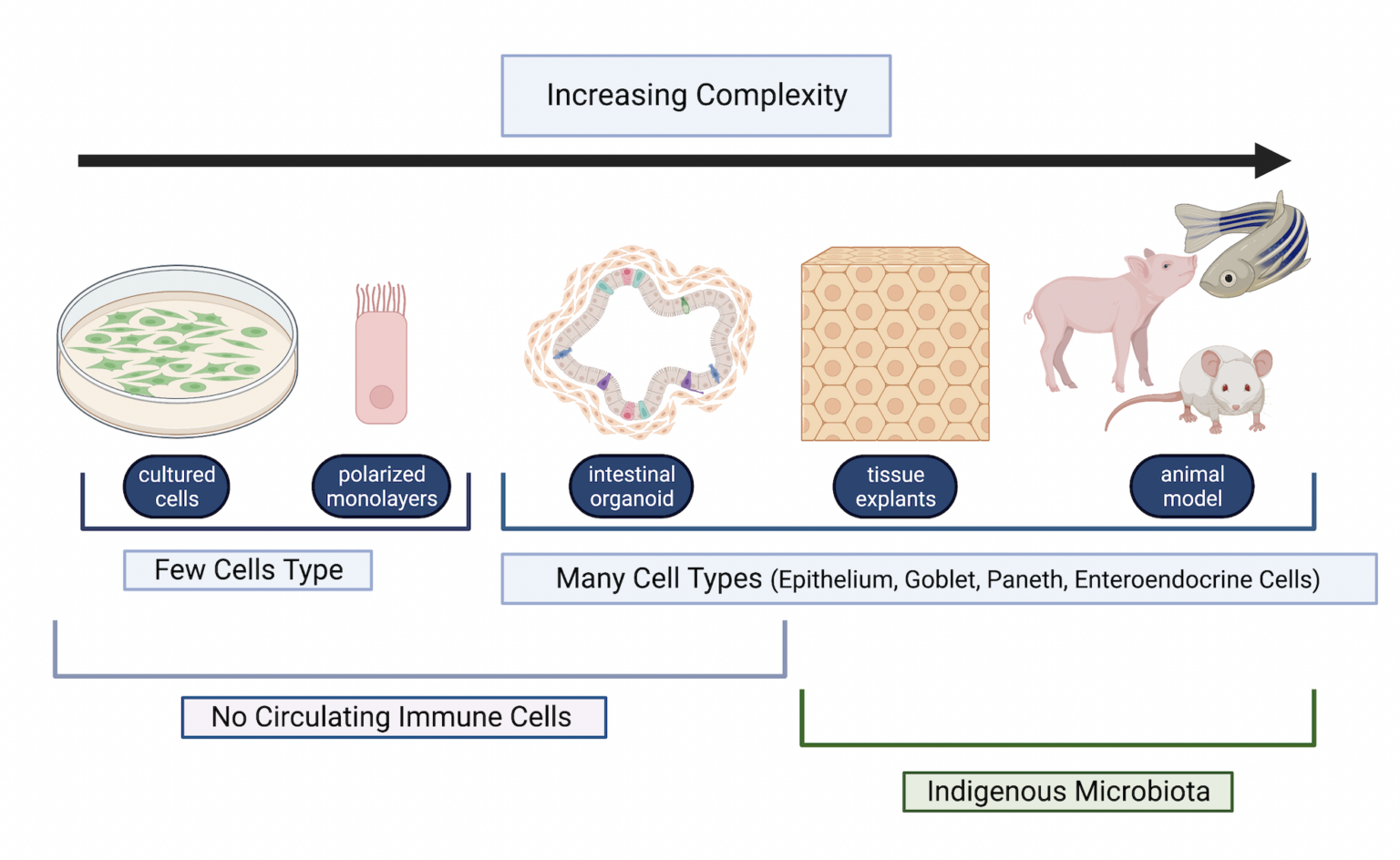
Figure 3. Comparison of model systems to study host-microbe interactions. The models are arranged in order of increasing complexity. Attributes of each model that recapitulate host-microbe interactions are indicated. (created in BioRender).
Expanding tools for genetic manipulation in both host and bacteria are also making zebrafish a more attractive model organism for studying the microbio. For instance, CRISPR-Cas genome editing is highly effective in zebrafish, making it possible to study the effects of host genetic mutations on the microbiota. Additionally, large-scale genetic screens have generated collections of zebrafish mutants, and collections of zebrafish-associated bacteria are well-defined.
Finally, zebrafish can also be used for “experimental evolution” experiments that would be difficult in mammals. For example, researchers at the University of Oregon have used zebrafish as a “living test tube” to rapidly evolve bacterial strains and discover bacterial features that regulate colonization of the host (Robinson, et al., 2021). Overall, zebrafish are a powerful model organism for studying the microbiota due to their highly-conserved systems, complex behavioral patterns, and expanding tools for genetic manipulation.
Disadvantages
Despite the many benefits of using zebrafish as a model for microbiota research, it is also important to keep disadvantages in mind when deciding which model is important for an experimental question. For example, the species-level composition of the intestinal microbiota in zebrafish is dramatically different from that of humans, which makes transplantation or humanization studies difficult to conduct (Stephens et al., 2016). The living environment of zebrafish, including temperature and diet, are also significantly different from that of humans, which can it more difficult to translate studies addressing the influence of diet on the microbiota. Finally, fish and mammalian lymphatic systems of fish are different, which can make it difficult to translate some studies of the immune system from fish to humans (Castranova et al., 2021). However, recent studies suggest that significant conservation exists between the mammalian and fish microbiota, just at the level of function rather than taxonomy (Xia et al., 2022).
As in other models, colonization dynamics are poorly defined for many species that typically colonize zebrafish. Though this can make it difficult to fully understand the interactions between the host and microbes, it remains an active area of new research (Chu, et al., 2021).
When comparing zebrafish to a traditional mammalian mouse model, it can be easy to focus on the fact that 70% of human genes have a homolog in fish but 99% of human genes have a homolog in mice. However, 82% of human disease-causing genes have a zebrafish counterpart! Thus, zebrafish represent an excellent first step for discovering host-microbe interactions in preclinical research, reducing the cost and complexity of subsequent studies in mammalian models.
Conclusion
Zebrafish are a valuable alternative model for studying host-microbe interactions due to their small size, experimental advantages, and significant homology to humans. Understanding host-microbe interactions is vital for understanding immune responses, longevity, and creating safer, more efficient drug therapeutics for countless conditions. Therefore, zebrafish are a great tool for accelerating improvement in human health, and at InVivo Biosystems we are doing our part, applying our zebrafish expertise to advance this field! Read Thaena’s Press Release to learn more.
References:
- Zheng, D., Liwinski, T. & Elinav, E. Interaction between microbiota and immunity in health and disease. Cell Res 30, 492–506 (2020). https://doi.org/10.1038/s41422-020-0332-7
- Parkin, K., Christophersen, C. T., Verhasselt, V., Cooper, M. N., & Martino, D. (2021). Risk Factors for Gut Dysbiosis in Early Life. Microorganisms, 9(10), 2066. https://doi.org/10.3390/microorganisms9102066
- A.I. Petra, S. Panagiotidou, E. Hatziagelaki, J.M. Stewart, P. Conti, T.C. Theoharides (2015). Gut-microbiota-brain Axis and its effect on neuropsychiatric disorders with suspected immune dysregulation, Clin. Therapeut., 37, pp. 984-995
- Carding, K. Verbeke, D.T. Vipond, B.M. Corfe, L.J. Owen (2015). Dysbiosis of the gut microbiota in disease, Microb. Ecol. Health Dis., 26, Article 26191
- Dikeocha, I.J., Al-Kabsi A. M., Miftahussurur, M. & Alshawsh (2022). Pharmacomicrobiomics: Influence of gut microbiota on drug and xenobiotic metabolism, FASEB, 36(6), https://doi.org/10.1096/fj.202101986R
- Zimmermann, M., Zimmermann-Kogadeeva, M., Wegmann, R. et al. Mapping human microbiome drug metabolism by gut bacteria and their genes. Nature 570, 462–467 (2019). https://doi.org/10.1038/s41586-019-1291-3
- Tan, F., Limbu, S. M., Qian, Y., Qiao, F., Du, Z. Y., & Zhang, M. (2019). The Responses of Germ-Free Zebrafish (Danio rerio) to Varying Bacterial Concentrations, Colonization Time Points, and Exposure Duration. Frontiers in microbiology, 10, 2156. https://doi.org/10.3389/fmicb.2019.02156
- MacRae, C. A., & Peterson, R. T. (2015). Zebrafish as tools for drug discovery. Nature reviews. Drug discovery, 14(10), 721–731. https://doi.org/10.1038/nrd4627
- Robinson, C. D., Sweeney, E. G., Ngo, J., Ma, E., Perkins, A., Smith, T. J., Fernandez, N. L., Waters, C. M., Remington, S. J., Bohannan, B. J. M., & Guillemin, K. (2021). Host-emitted amino acid cues regulate bacterial chemokinesis to enhance colonization. Cell host & microbe, 29(8), 1221–1234.e8. https://doi.org/10.1016/j.chom.2021.06.003
- Flores, E. M., Nguyen, A. T., Odem, M. A., Eisenhoffer, G. T., & Krachler, A. M. (2020). The zebrafish as a model for gastrointestinal tract-microbe interactions. Cellular microbiology, 22(3), e13152. https://doi.org/10.1111/cmi.13152
- Levraud, JP., Rawls, J.F. & Clatworthy, A.E. Using zebrafish to understand reciprocal interactions between the nervous and immune systems and the microbial world. J Neuroinflammation 19, 170 (2022). https://doi.org/10.1186/s12974-022-02506-x
- Chu, N. D., Crothers, J. W., Nguyen, L. T. T., Kearney, S. M., Smith, M. B., Kassam, Z., Collins, C., Xavier, R., Moses, P. L., & Alm, E. J. (2021). Dynamic Colonization of Microbes and Their Functions after Fecal Microbiota Transplantation for Inflammatory Bowel Disease. mBio, 12(4), e0097521. https://doi.org/10.1128/mBio.00975-21
- Stephens, W. Z., Burns, A. R., Stagaman, K., Wong, S., Rawls, J. F., Guillemin, K., & Bohannan, B. J. (2016). The composition of the zebrafish intestinal microbial community varies across development. The ISME journal, 10(3), 644–654. https://doi.org/10.1038/ismej.2015.140
- Castranova, D., Samasa, B., Venero Galanternik, M., Jung, H. M., Pham, V. N., & Weinstein, B. M. (2021). Live Imaging of Intracranial Lymphatics in the Zebrafish. Circulation research, 128(1), 42–58. https://doi.org/10.1161/CIRCRESAHA.120.317372
- Xia, H., Chen, H., Cheng, X. et al. Zebrafish: an efficient vertebrate model for understanding role of gut microbiota. Mol Med 28, 161 (2022). https://doi.org/10.1186/s10020-022-00579-1
- Appleton J. (2018). The Gut-Brain Axis: Influence of Microbiota on Mood and Mental Health. Integrative medicine (Encinitas, Calif.), 17(4), 28–32.
- Maseda, D., & Ricciotti, E. (2020). NSAID-Gut Microbiota Interactions. Frontiers in pharmacology, 11, 1153. https://doi.org/10.3389/fphar.2020.01153
Unlock the power of drug toxicity testing in zebrafish for precise and efficient chemical evaluation. Contact us today to access our cutting-edge service and make informed decisions for a safer future.




