Summary
New ‘humanized models’ have the potential to revolutionize the study of clinical diseases, creating models with higher validity than more traditional models. This in turn has the power to expedite getting results from the bench to the bedside. In this article we will discuss the current state of animal models and the future that humanized models make possible; as an example, we will specifically focus on the models of Alzheimer’s Disease (AD).
Research has long relied on animal models to recapitulate human diseases, but new gene editing techniques are enabling new models to be developed: ‘humanized models‘. These are models which have had human genes inserted into their genome, enabling in vivo analysis, and helping to move the field towards a more personalized approach to medicine.
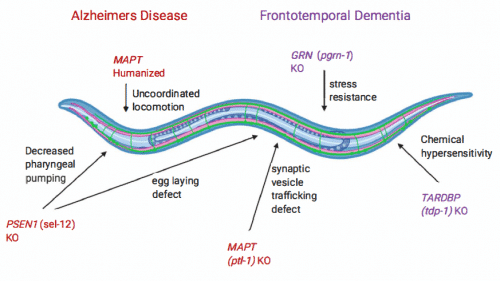
Figure 1. C. elegans as a transgenic model for studying Alzheimer’s Disease (AD) and Frontotemporal Dementia (FD).
Humanized models have been referred to as the, “new generation of models” because they bridge the gap between traditional animal models and human research (Figure 2) (Viteket al., 2021). Take, for instance, research into Alzheimer’s Disease (AD); there is growing awareness that there are many factors involved in AD (age, sex, race, ethnicity, genetics, comorbitidites, etc.), which result in mixed pathologies. Currently, however, models are failing to reflect the entire biology of this disorder, making research into treatment options difficult and as a result AD is still untreatable. Recently, humanization technology has allowed researchers to successfully create models that test the hallmarks of AD alongside common comorbidities of AD (Meakin et al., 2020). One example of this is a model that tests the link between endothelial dysfunction (a vascular disease commonly associated with obesity, and diabetes) with AD. In this study, the human amyloid precursor protein (APP) gene was introduced to mice, who were fed a diet to mimic obesity level. These mice were then treated with an APP-cleaving inhibitor (BACE1). The results suggest that the BACE1 inhibitors could be a novel therapy for vascular dysfunction (which often occurs early in AD progression, sometimes diagnosed decades before AD). More studies need to be done on the co-occurrence of vascular disease and AD, as it is an intriguing avenue of research.
Similarly, a limitation to traditional models are their inability to mimic human immune responses. Humanization may offer a more predictive way of modeling, as humanized mice models with APOE (a protein whose subtype is correlated with AD) backgrounds have clearly shown that increased behavioral and pathological deficits that mimic AD pathogenesis (Tai et al., 2017). Therefore, these models provide more translatable systems, as the introduction of human genes lessens the suppression and/or activation of the animal’s immune system (Viteket al., 2021).
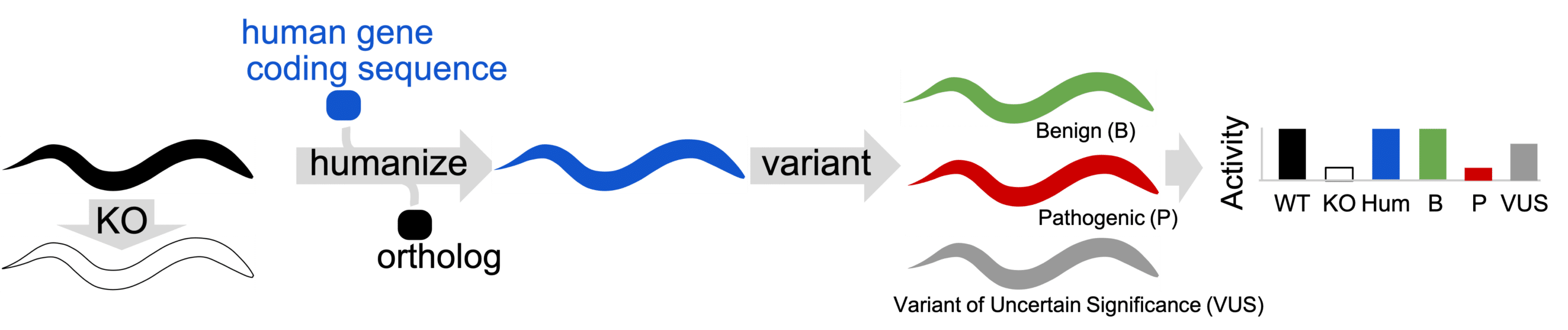
Figure 2: At InVivo Biosystems we use the Gene-swap humanization technique: in this technique an Ortholog gene is removed from animal (KO) to determine if defective phenotype is present. Then, the Ortholog is replaced with human coding sequence to create gene-humanized animal. Variants are installed in gene-humanized locus create Patient Avatars. Humanized animals in Patient Avatar format are phenotyped for activity defects.
Lastly, humanization can aid in Alzheimer’s research by helping to expand the field of research, creating new genetic models of a largely under-studied type of AD: late-onset Alzheimer’s Disease (LOAD). Alzheimer’s is often categorized into two main types: early-onset Alzheimer’s Disease (people <65), and late-onset Alzheimer’s Disease (people 65 and older). While the vast majority of AD cases are the late-onset type (>95%), most research has been conducted on the early-onset type. This can be attributed to the fact that most early-onset cases are Familial Alzheimer’s disease, and have been linked to specific single-gene mutations (NIH, 2019b). Researchers are hopeful that by adopting humanized models, more specific models can be developed, and at a faster rate, which can help the high failure rate (99.6%) of AD drugs. Expanding the use of other, non-mammalian models, such as C. elegans and zebrafish, can be particularly helpful in accelerating the rate of disease-specific research as these are models which have a shorter life-span than mice.
Non-mammalian models can act as excellent models for disease-specific research as their genome is surprisingly homologous to the human genome (of all disease causing genes in humans 65% have a C. elegans ortholog, and 82% have a zebrafish ortholog) (Baumeister & Ge, 2002; Howe et al., 2013). These non-mammalian models are also extremely well-suited for genetic manipulation thanks to their simplicity and optical transparency. InVivo Biosystems is at the forefront of using non-mammalian models and new technologies, such as CRISPR, to better understand diseases. We have already developed successful humanized models of AD (C. elegans) and Epilepsy (C. elegans and zebrafish), which show some of the hallmarks of neurodegeneration seen in these disorders.
Moreover, the use of non-mammalian models benefit the research field overall, as it will enable more research to be conducted since mice models are not always accessible. Mice colonies require a sizable amount of space, and considerable funds, whereas non-mammalian models take a fraction of these resources. Thus, this new generation of humanized models can help to make the scientific community more diverse and inclusive.
The past century has seen a dramatic rise in genetic studies; breeding animals specifically for genetic studies began in 1902, and now genetic modification accounts for nearly half (49%) of animal procedure in the UK (Ericsson, Crim & Franklin, 2013; Home Office, 2019). While most of the models currently being used are mouse models, there have been great successes with fish (such as zebrafish), and c. elegans among others. Indeed, fish make up 12% of the remaining 13% of genetically modified animals which were not mice in UK (these numbers are recorded due to the Animals Scientific Procedures Act 1986, and, as a result, do not include invertebrates such as C. elegans, or Drosophila melanogaster).
This rapid adoption of genetic studies has changed the landscape of research, allowing the research community to make great strides in creating a more personalized version of medicine. Thanks to the CRISPR/Cas9 technology, the next generation of disease-focused models can be developed faster, and have a more targeted approach. These new models should consist of more than one type of animal model, as to use the advantages of each organism, and comorbidities/environmental factors should be considered when testing them. All in all, these models present a promising future for finding new therapeutics for some of humankind’s most prevalent and debilitating illnesses.
References
- Baumeister, R., & Ge, L. (2002). The worm in us – Caenorhabditis elegans as a model of human disease. Trends in biotechnology, 20(4), 147-148. https://doi.org/10.1016/s0167-7799(01)01925-4
- Ericsson, A.C., Crim, M.J. & Franklin, C.L. (2013). A Brief History of Animal Modeling. Missouri Medicine. 110(3): 201-205. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3979591/
- Howe, K., Clark, M. D., Torroja, C. F., Torrance, J., Berthelot, C., Muffato, M., Collins, J. E., Humphray, S., McLaren, K., Matthews, L., McLaren, S., Sealy, I., Caccamo, M., Churcher, C., Scott, C., Barrett, J. C., Koch, R., Rauch, G. J., White, S., Chow, W., … Stemple, D. L. (2013). The zebrafish reference genome sequence and its relationship to the human genome. Nature, 496(7446), 498-503. https://doi.org/10.1038/nature12111
- Meakin, P. J., Coull, B. M., Tuharska, Z., McCaffery, C., Akoumianakis, I., Antoniades, C., Brown, J., Griffin, K. J., Platt, F., Ozber, C. H., Yuldasheva, N. Y., Makava, N., Skromna, A., Prescott, A., McNeilly, A. D., Siddiqui, M., Palmer, C. N., Khan, F., & Ashford, M. L. (2020). Elevated circulating amyloid concentrations in obesity and diabetes promote vascular dysfunction. The Journal of clinical investigation, 130(8), 4104-4117. https://doi.org/10.1172/JCI122237
- Neff, E.P. Animal models of Alzheimer’s disease embrace diversity. Lab Anim 48, 255-259 (2019). https://doi.org/10.1038/s41684-019-0377-8
- Tai, L. M., Balu, D., Avila-Munoz, E., Abdullah, L., Thomas, R., Collins, N., Valencia-Olvera, A. C., & LaDu, M. J. (2017). EFAD transgenic mice as a human APOE relevant preclinical model of Alzheimer’s disease. Journal of lipid research, 58(9), 1733-1755. https://doi.org/10.1194/jlr.R076315
- https://www.understandinganimalresearch.org.uk/files/8615/9955/8508/annual-statistics-scientific-procedures-living-animals-2019.pdf