Summary
A recent publication by Crouzier et al. emphasized the utility of zebrafish as a model for investigating rare diseases – enabling better understanding of their underlying mechanisms, and the development of therapeutics to treat these diseases. In this article we will discuss Crouzier et al.’s publication, and InVivo Biosystems’ ongoing work to help identify and study variants of interest.
Rare diseases are defined as conditions which affect less than 1 in 2,000 people, and while individually these diseases only impact a small percentage of the population, there are over 800 rare diseases worldwide which collectively affect more than 400 million people (Crouzier et al., 2021). Thus, rare diseases pose a huge strain, not only on individual people, but on their communities as well [Image 1]. As previously mentioned, however, the rare disease category consists of over 800 diseases – thus, it is particularly difficult to study rare diseases because the number of patients any one disease affects is so relatively small that the traditional methods used in disease research and drug discovery are impractically time-consuming and expensive. Consequently, alternative, robust, models are needed for rare disease research. In their recent publication, Crouzier et al., make the argument for the small tropical zebrafish (Danio Rerio) to be the new model of choice.
Image 1. How prominent rare diseases are when considered collectively.
Zebrafish have some notable benefits for rare disease research: a short lifespan and large brood sizes, a fully sequenced genome with homologues for over 84% of disease-associated genes in humans, and organ systems that are remarkably similar to humans’.
Crouzier et al. specifically called out the impact that the advent of genetic tools, such as TALENs and CRISPR/Cas9, has had on zebrafish’s usefulness as a model.
“These methods [TALENS, CRISPR/Cas9] have been optimized in zebrafish and they have proven to be fast and cost-effective” – Crouzier et al. (2021)
These tools have led to zebrafish being extensively used in targeted drug screening – as these tools enable the creation of humanized fish [Image 2]. At InVivo Biosystems we generate humanized zebrafish models for R&D researchers, academics, and pharmaceutical companies alike. Our Human Avatars have enabled unknown variants of significance (VUS) to be identified as benign and pathogenic, and in collaboration with the Undiagnosed Disease Network (UDN) our team is leading the way to better understanding of disease mutations.
Image 2. Transgenic zebrafish can be generated and are able to act as Humanized models for Rare Diseases
Example of InVivo Biosystems’ Work
Dr Sahar Da’as is a long time collaborator with InVivo Biosystems. In 2014 she established the first zebrafish facility within Qatar and now leads the Sidra Medicine Zebrafish Facility. Dr Da’as models mutations seen in the clinic, such as rare hereditary pediatric diseases – often utilizing crispants (loss of function first generation larvae) that our team at InVivo Biosystems generates as a fast way to assess phenotypes. Thus, zebrafish models are not only allowing Dr. Da’as to validate and characterize the genetic variants found in Qatar’s clinics, but more broadly are pushing the medical community towards a personalized approach.
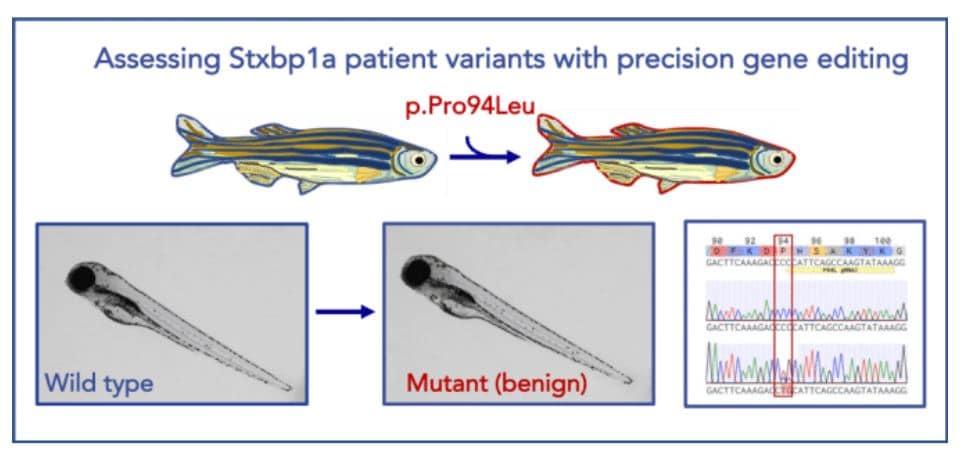
Image 3. Illustration of a successful precise humanized point mutation of Stxbp1a in zebrafish. STXBP1a is a highly conserved zebrafish ortholog of human STXBP1 (87% identity). Using CRISPR/Cas9 technology, we were able to precisely generate a benign patient mutation at the conserved amino acid residue (CCC>CTG, p.P94L).Learn more about our humanization capabilities
References
- Crouzier, L., Richard, E. M., Sourbron, J., Lagae, L., Maurice, T., & Delprat, B. (2021). Use of Zebrafish Models to Boost Research in Rare Genetic Diseases. International journal of molecular sciences, 22(24), 13356. https://doi.org/10.3390/ijms222413356
Take a step towards more ethical and efficient toxicity testing with our zebrafish toxicity testing service. By leveraging the remarkable attributes of zebrafish as a model organism, we offer a cutting-edge solution for evaluating the safety of chemicals and substances.



