Epilepsy Genetics
1 out of 100 persons is living with active epilepsy (Zack and Kobau 2017, WHO 2018). For the subset that can be pinned down to having a genetic cause, there are about 70 or so genes involved in causing the illness (Lindy 2018). Until recently, the frequency of a gene’s association with epilepsy remained unclear. The GeneDX study by the Lindy team is helping bring clarity by analyzing the genetic underpinnings in 8565 patients with active epilepsy.
Positive Cases
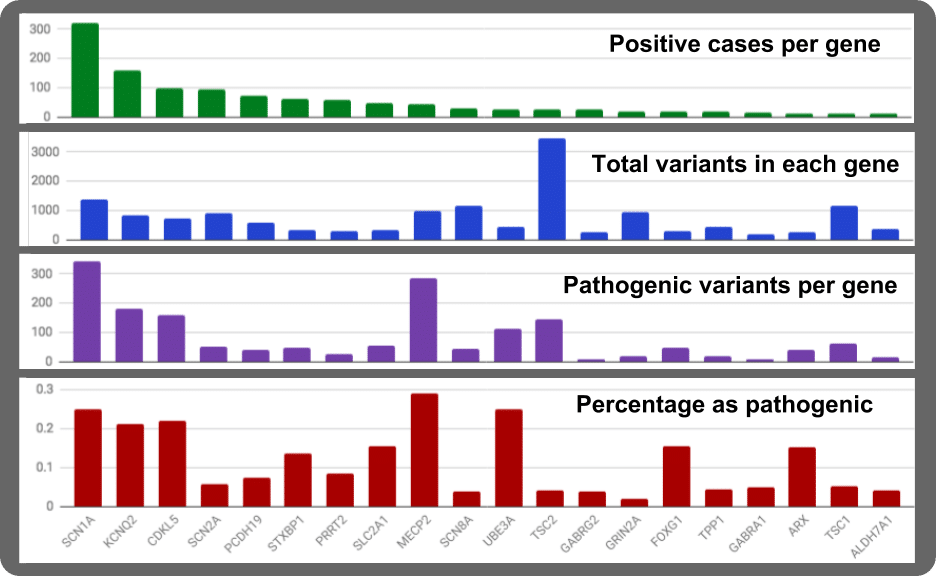
From the GeneDX study, we can plot the rank of the top 20 genes in epilepsy (green graph at top of the figure). The SCN1A gene is by far the major source gene for genetically-associated epilepsies. 27% of all the pathogenic variants in the top 20 epilepsy genes are in SCN1A and 24% in all 70 known epilepsy genes. For instance, 322 positive cases occur in SCN1A out of a total of 1181 positive case in the top 20 genes. Applying the data set to the larger populations the 1181 cases of 8565 (13%) as genetically caused, suggest close to 1 of 1000 persons are living with gene-induced epilepsy. For SCN1A, the population estimate is 78,000 individuals in USA or 1.8 Million worldwide living with pathogenic lesions in their SCN1A gene.
Total Variants
Another way to look at the rank is to ask how many variants occur in each gene (Blue graph). Gathering data from NCBI’s Variant Viewer (https://www.ncbi.nlm.nih.gov/variation/view) There is a large difference in numbers of observed variant per gene. For example, in two genes of similar protein size (SCN1A and TSC2), there is a 2.5 fold difference in their relative numbers of variants.
Pathogenic Variants
In another aspect derived from Variant Viewer, we can look at numbers of variants known to be pathogenic. For the top 20 epilepsy genes, SCN1A comes out on top again but the next gene with high levels of pathogenic variants is MECP2 (purple graph).
Pathogenic vs. Total
Finally, when we look at the ratio of pathogenic to total variants, we see some interesting findings. MECP2 has major sensitivity by having a high pathogenic variant load. Similarly, UBE3 also harbors a high pathogenic variant ratio. Another gene jumping up for sensitivity is the expected SCN1A gene. But now we also get KCNQ2, CDKL5, STXBP1, SLC2A1, FOXG1, and ARX as variation-sensitive genes.
Favorite Genes and an Anomaly
A gene for which I hold high passion is STXBP1. This gene is ranked #6 as likely genetic cause of epilepsy. STXBP1 has 47 variants of 349 as pathogenic (13%). Another gene on our list for humanization development is KCNQ2. This gene is at the #2 position for being a frequent cause of epilepsy. KCNQ2 has 855 variants of which 181 are pathogenic so it has a pathogenicity load of 21%. The TSC2 gene has a strange result. There are 145 pathogenic variants out of 3445 total which gives a 4% pathogenicity penetration.
Why are there so many non-pathogenic variants for this gene? Perhaps this gene is highly flexible and can tolerate a high level of variant load. There is a slightly higher proportion of synonymous variants at 27%. Population frequency in the measured variant pool exceeds 0.0001 for only 12% of the variants. So, most of the TSC2 alleles are rare. My other suspicion is TSC2 is like BRCA1 gene. There has been the very large size of researchers studying each of these genes. This leads to more patients being examined for variants in TSC2 and BRCA1. The result, these two genes have attained a higher sampling of the variant diversity occurring in the general population.
Citations
- Zack MM and Kobau R. National and State Estimates of the Numbers of Adults and Children with Active Epilepsy – United States, 2015. CDC: MMWR Morb Mortal Wkly Rep. 2017 Aug 11;66(31):821-825. doi: 10.15585/mmwr.mm6631a1. https://www.ncbi.nlm.nih.gov/pubmed/28796763
- Epilepsy (World Health Organization, access 8/1/2018) http://www.who.int/mediacentre/factsheets/fs999/en/
- Lindy AS et al. Diagnostic outcomes for genetic testing of 70 genes in 8565 patients with epilepsy and neurodevelopmental disorders. Epilepsia. 2018 May;59(5):1062-1071. doi: 10.1111/epi.14074. Epub 2018 Apr 14. https://www.ncbi.nlm.nih.gov/pubmed/29655203